elegans β-amyloid aggregation model in a PINK-1-dependent manner. Importantly, we found that treatment with two compounds, which we named PS83 and PS106 (more commonly known as sertraline) reduced neurodegenerative disease phenotypes, including delaying paralysis in a C. Several of the compounds also reduced ATP production, oxygen consumption, mitochondrial mass, and/or mitochondrial membrane potential. We obtained eight hits that increased mitochondrial fragmentation and autophagosome formation. Toward this end, we screened ~ 45,000 small molecules using a high-throughput, whole-organism, phenotypic screen that monitored accumulation of PINK-1 protein, a key event in mitophagic activation, in a Caenorhabditis elegans strain carrying a Ppink-1::PINK-1::GFP reporter. This has led to the hypothesis that pharmacological stimulation of mitophagy may be a feasible approach to combat neurodegeneration. Accumulation of defective mitochondria and impaired mitophagy have been widely implicated in many neurodegenerative diseases, and loss-of-function mutations of PINK1 and Parkin, two key regulators of mitophagy, are amongst the most common causes of heritable parkinsonism. Again, this is quite obvious to the eye but I can’t find a way to identify it with CellProfiler.Macroautophagic recycling of dysfunctional mitochondria, known as mitophagy, is essential for mitochondrial homeostasis and cell viability. In the image “H07_90_DAPI_fluorescencePattern.tif”, the fluorescence pattern is different from the negative control – there is an accumulation of fluorescence in the anterior part of the worm. I have also tried the module IdentifyDeadWorms using the WormBinary image obtained from UntangleWorms #15 and it doesn’t quite work with these images.ī) Distinguish a particular fluorescence pattern: I have inspected the measurements produced by MeasureTexture and MeasureGranularity modules and I haven’t been able to find a parameter that is clearly different in the dead worm images. 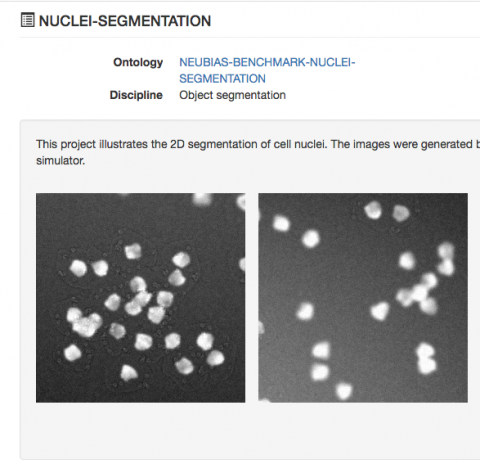
“H07_90_DAPI_fluorescencePattern.tif” image), so I can’t really use this parameter. The fluorescence ( MeanIntensity) is slightly lower in dead worms, but other compounds also cause a reduction in fluorescence without killing the worms (e.g. It seems that the area, compactness, majoraxislength, and perimeter are slighlty higher in dead worms compared to the negative controls (see distribution plots attached), but the difference is not big enough to be able to establish a clear threshold. While these images look quite different than the negative controls (“negCtrl”) to the eye, I would like to be able to distinguish them using CellProfiler. Some compounds kill the worms (labeled as “deadWorms” in example images). The data I get out of this pipeline is already very useful but I am struggling to find a way to do the following:Ī) Distinguish dead worms from alive worms: Count the number of worm progeny objects.Measure fluorescence intensity, texture, and granularity of worm objects in fluorescence images.Using CellProfiler and the WormToolbox I have managed to put together a pipeline that does the following: elegans treated with different compounds that alter the worm fluorescence intensity and/or pattern. I am analysing fluorescence images from C.
0 Comments
Leave a Reply. |

 RSS Feed
RSS Feed
